DARPA-funded ‘body on a chip’ microfluidic system could revolutionize drug evaluation
March 19, 2018
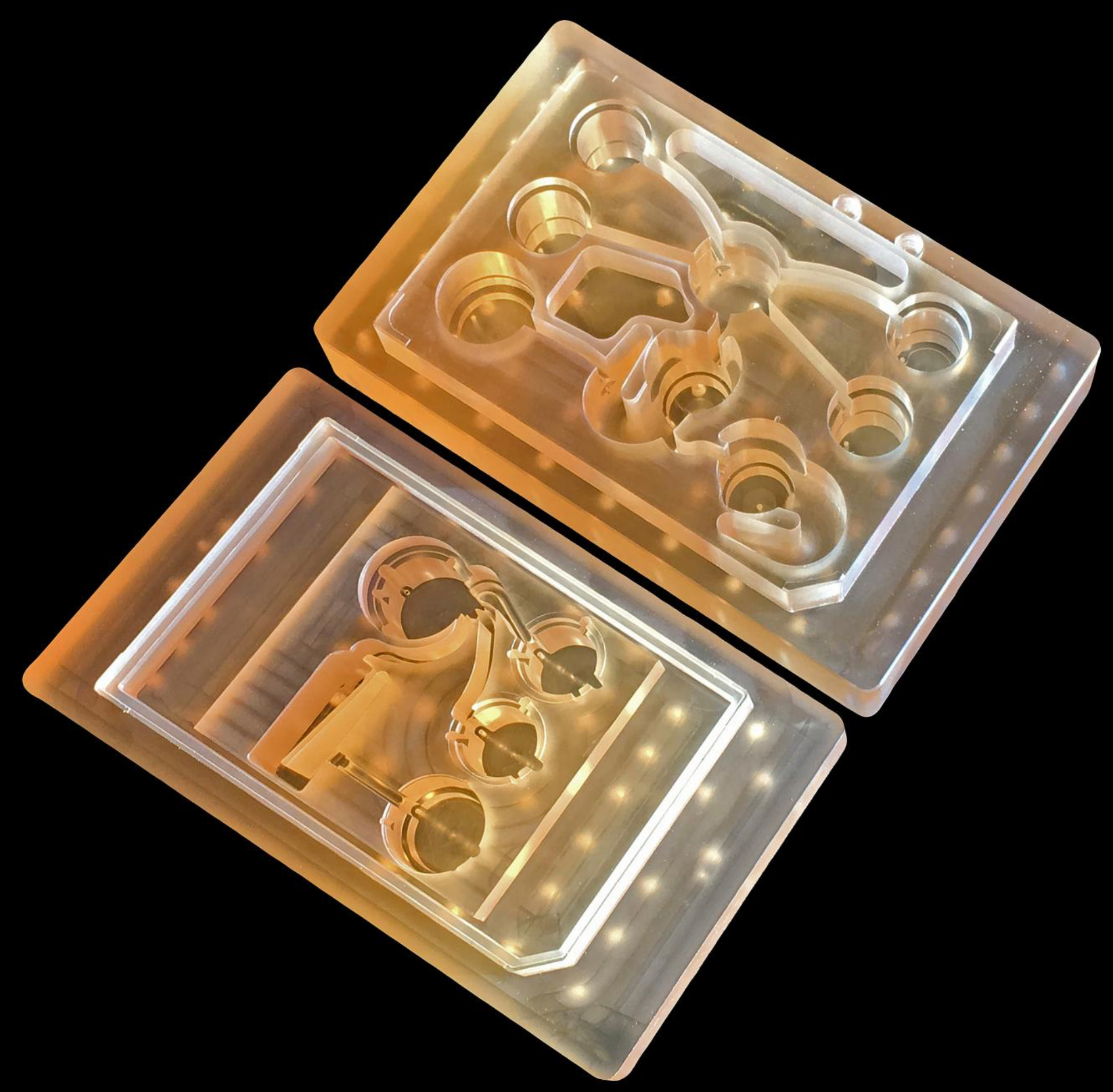

To measure the effects of drugs on different parts of the body, MIT’s new “Physiome-on-a-chip” microfluidic platform can connect 3D tissues from up to 10 “organs on chips” — allowing researchers to accurately replicate human-organ interactions for weeks at a time. (credit: Felice Frankel)
MIT bioengineers have developed a new microfluidic platform technology that could be used to evaluate new drugs and detect possible side effects before the drugs are tested in humans.
The microfluidic platform can connect 3D tissues from up to 10 organs. Replacing animal testing, it can accurately replicate human-organ interactions for weeks at a time and can allow for measuring the effects of drugs on different parts of the body, according to the engineers. For example, the system could reveal whether a drug that is intended to treat one organ will have adverse effects on another.
Physiome on a chip. The new technology was originally funded in 2012 by the Defense Advanced Research Projects Agency (DARPA) Microphysiological Systems (MPS) program (see “DARPA and NIH to fund ‘human body on a chip’ research”). The goal of the $32 million program was to model potential drug effects more accurately and rapidly.
Linda Griffith, PhD, the MIT School of Engineering Professor of Teaching Innovation, a professor of biological engineering and mechanical engineering, and her colleagues decided to pursue a technology that they call a “physiome on a chip.” Griffith is one of the senior authors of a paper on the study, which appears in the open-access Nature journal Scientific Reports.
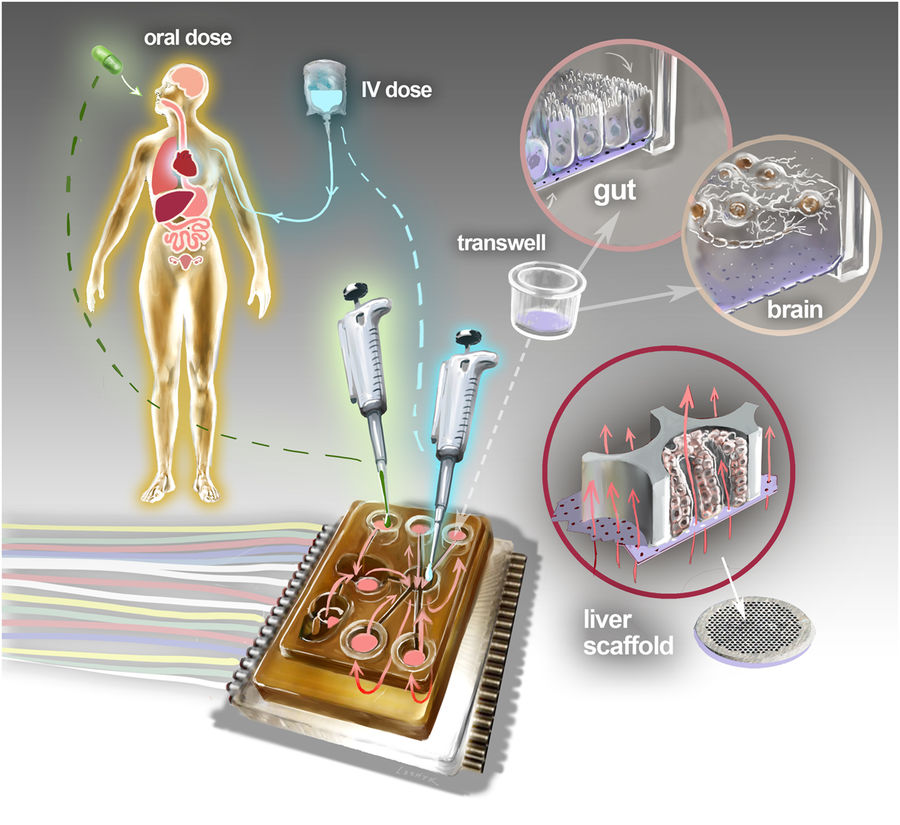
Schematic overview of “Physiome-on-a-chip” approach, using bioengineered devices that nurture many interconnected 3D “microphysiological systems” (MPSs), aka “organs-on-chips.” MPSs are in vitro (lab) models that capture facets of in vivo (live) organ function. They represent specified functional behaviors of each organ of interest, such as gut, brain, and liver, as shown here. MPSs are designed to capture essential features of in vivo physiology that is based on quantitative systems models tailored for individual applications, such as drug fate or disease modeling. (illustration credit: Victor O. Leshyk)
To achieve this, the researchers needed to develop a platform that would allow tissues to grow and interact with each other. They also needed to develop engineered tissue that would accurately mimic the functions of human organs. Before this project was launched, no one had succeeded in connecting more than a few different tissue types on a platform. And most researchers working on this kind of chip were working with closed microfluidic systems, which allow fluid to flow in and out but do not offer an easy way to manipulate what is happening inside the chip. These systems also require awkward external pumps.
The MIT team decided to create an open system, making it easier to manipulate the system and remove samples for analysis. Their system was adapted from technology they previously developed and commercialized through U.K.-based CN BioInnovations*. It also incorporates several on-board pumps that can control the flow of liquid between the “organs,” replicating the circulation of blood, immune cells, and proteins through the human body. The pumps also allow larger tissues (for example, tumors within an organ) to be evaluated.
Complex interactions with 1 or 2 million cells in 10 organs. The researchers created three versions of their system, linking up to 10 organ types: liver, lung, gut, endometrium, brain, heart, pancreas, kidney, skin, and skeletal muscle. Each “organ” consists of clusters of 1 million to 2 million cells.
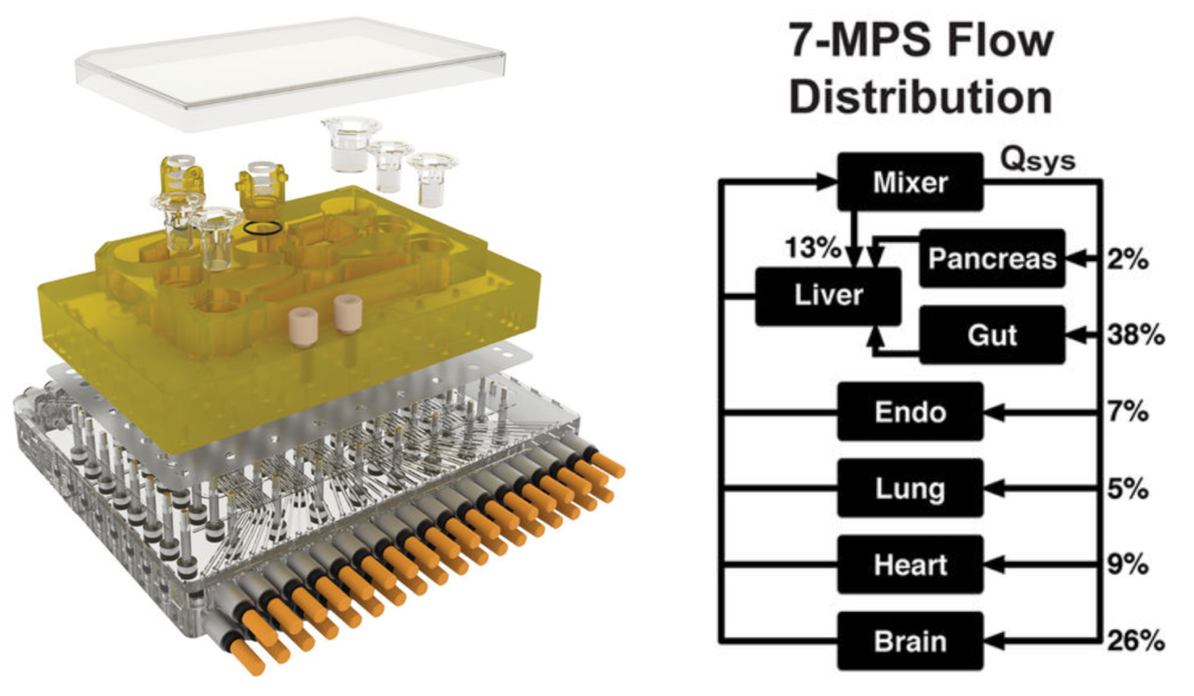
MPS platform and flow partitioning. (Left) Exploded rendering of the 7-MPS platform. Rigid plates of polysulfone (yellow) and acrylic (clear) sandwich an elastomeric (rubbery) membrane to form a pumping manifold with integrated fluid channels. Channels interface to the top side of the polysulfone plate (yellow) to deliver fluid to each MPS compartment (such as brain) in a defined manner. Fluid return occurs passively via spillway channels machined into the top plate. (Right) Flow partitioning mirrors physiological cardiac output for 7-way platforms. A 10-MPS platform adds muscle, skin, and kidney flow. (credit: Collin D. Edington et al./Scientific Reports)
These tissues don’t replicate the entire organ, but they do perform many of its important functions. Significantly, most of the tissues come directly from patient samples rather than from cell lines that have been developed for lab use. These “primary cells” are more difficult to work with, but offer a more representative model of organ function, Griffith says.
Using this system, the researchers showed that they could deliver a drug to the gastrointestinal tissue, mimicking oral ingestion of a drug, and then observe as the drug was transported to other tissues and metabolized. They could measure where the drugs went, the effects of the drugs on different tissues, and how the drugs were broken down.
Replacing animal testing. The new microfluidic platform technology is superior to animal testing, according to Griffith:
- Preclinical testing in animals can offer information about a drug’s safety and effectiveness before human testing begins, but those tests may not reveal potential side effects.
- Drugs that work in animals often fail in human trials.
- “Some of these effects are really hard to predict from animal models because the situations that lead to them are idiosyncratic. With our chip, you can distribute a drug and then look for the effects on other tissues and measure the exposure and how it is metabolized.”
- These chips could also be used to evaluate antibody drugs and other immunotherapies, which are difficult to test thoroughly in animals because the treatments are only designed to interact with the human immune system.
Modeling Parkinson’s and metastasizing tumors. Griffith believes that the most immediate applications for this technology involve modeling two to four specific organs. Her lab is now developing a model system for Parkinson’s disease that includes brain, liver, and gastrointestinal tissue, which she plans to use to investigate the hypothesis that bacteria found in the gut can influence the development of Parkinson’s disease. Other applications the lab is investigating include modeling tumors that metastasize to other parts of the body.
“An advantage of our platform is that we can scale it up or down and accommodate a lot of different configurations,” Griffith says. “I think the field is going to go through a transition where we start to get more information out of a three-organ or four-organ system, and it will start to become cost-competitive because the information you’re getting is so much more valuable.”
The research was funded by the U.S. Army Research Office and DARPA.
* Co-author David Hughes is an employee of CN BioInnovations, the commercial vendor for the Liverchip. Linda Griffith and Steve Tannenbaum receive patent royalties from the Liverchip.
Abstract of Interconnected Microphysiological Systems for Quantitative Biology and Pharmacology Studies
Microphysiological systems (MPSs) are in vitro models that capture facets of in vivo organ function through use of specialized culture microenvironments, including 3D matrices and microperfusion. Here, we report an approach to co-culture multiple different MPSs linked together physiologically on re-useable, open-system microfluidic platforms that are compatible with the quantitative study of a range of compounds, including lipophilic drugs. We describe three different platform designs – “4-way”, “7-way”, and “10-way” – each accommodating a mixing chamber and up to 4, 7, or 10 MPSs. Platforms accommodate multiple different MPS flow configurations, each with internal re-circulation to enhance molecular exchange, and feature on-board pneumatically-driven pumps with independently programmable flow rates to provide precise control over both intra- and inter-MPS flow partitioning and drug distribution. We first developed a 4-MPS system, showing accurate prediction of secreted liver protein distribution and 2-week maintenance of phenotypic markers. We then developed 7-MPS and 10-MPS platforms, demonstrating reliable, robust operation and maintenance of MPS phenotypic function for 3 weeks (7-way) and 4 weeks (10-way) of continuous interaction, as well as PK analysis of diclofenac metabolism. This study illustrates several generalizable design and operational principles for implementing multi-MPS “physiome-on-a-chip” approaches in drug discovery.
