NIH Human Microbiome Project defines normal bacterial makeup of the body
June 15, 2012
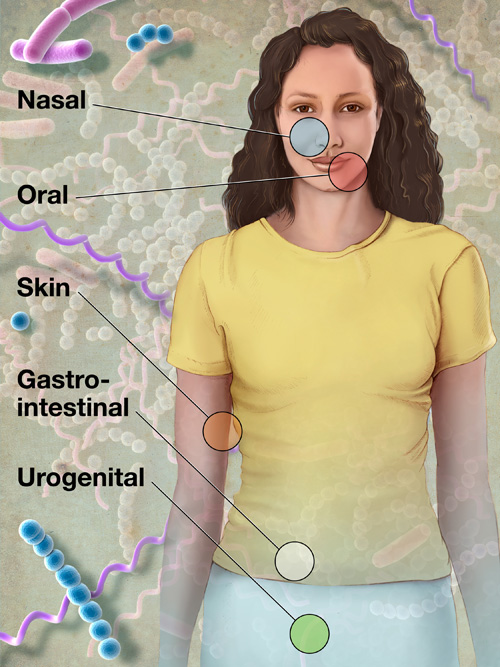
This illustration shows the body sites that will be sampled from volunteers for the Human Microbiome Project, part of the National Institutes of Health's Roadmap for Medical Research (credit:: NIH Medical Arts and Printing)
Some 200 members of the Human Microbiome Project (HMP) Consortium from nearly 80 universities and scientific institutions, organized by the National Institutes of Health, have mapped the normal microbial makeup of healthy humans, producing numerous insights and even a few surprises.
The report on on their five years of research was published Thusday June 14, 2012, in a series of coordinated scientific reports in Nature the PLoS.
Researchers found, for example, that nearly everyone routinely carries pathogens, microorganisms known to cause illnesses.
In healthy individuals, however, pathogens cause no disease; they simply coexist with their host and the rest of the human microbiome, the collection of all microorganisms living in the human body.
Researchers must now figure out why some pathogens turn deadly and under what conditions, likely revising current concepts of how microorganisms cause disease.
“Like 15th century explorers describing the outline of a new continent, HMP researchers employed a new technological strategy to define, for the first time, the normal microbial makeup of the human body,” said NIH Director Francis S. Collins, M.D., Ph.D.
“HMP created a remarkable reference database by using genome sequencing techniques to detect microbes in healthy volunteers. This lays the foundation for accelerating infectious disease research previously impossible without this community resource.”
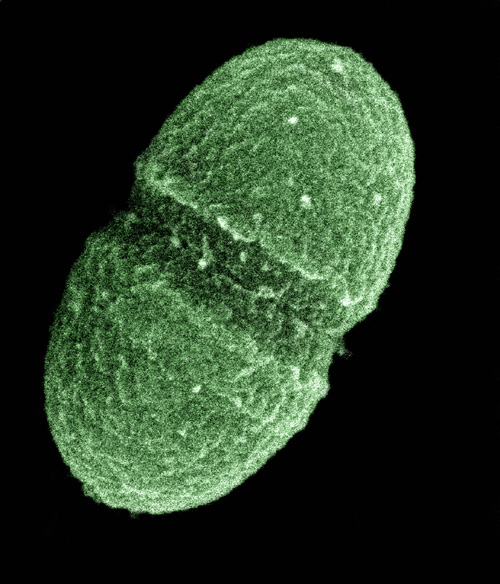
The bacterium, Enterococcus faecalis, which lives in the human gut, is just one type of microbe that will be studied as part of NIH's Human Microbiome Project (credit: United States Department of Agriculture)
To define the normal human microbiome, HMP researchers sampled 242 healthy U.S. volunteers (129 male, 113 female), collecting tissues from 15 body sites in men and 18 body sites in women.
Researchers collected up to three samples from each volunteer at sites such as the mouth, nose, skin (two behind each ear and each inner elbow), and lower intestine (stool), and three vaginal sites in women; each body site can be inhabited by organisms as different as those in the Amazon Rainforest and the Sahara Desert.
Historically, doctors studied microorganisms in their patients by isolating pathogens and growing them in culture. This painstaking process typically identifies only a few microbial species, as they are hard to grow in the laboratory. In HMP, researchers purified all human and microbial DNA in each of more than 5,000 samples and analyzed them with DNA sequencing machines.
Using computers, researchers sorted through the 3.5 terabases of genome sequence data to identify specific genetic signals found only in bacteria — the variable genes of bacterial ribosomal RNA called 16S rRNA. Bacterial ribosomal RNA helps form the cellular structures that manufacture protein and can identify the presence of different microbial species.
Focusing on this microbial signature allowed HMP researchers to ignore the human genome sequences and analyze only the bacterial DNA. In addition, metagenomic sequencing, or sequencing all of the DNA in a microbial community, allowed the researchers to study the metabolic capabilities encoded in the genes of these microbial communities.
“Recently developed genome sequencing methods now provide a powerful lens for looking at the human microbiome,” said Eric D. Green, M.D., Ph.D., director of the National Human Genome Research Institute, which managed HMP for NIH. “The astonishing drop in the cost of sequencing DNA has made possible the kind of large survey performed by the Human Microbiome Project.”
Where doctors had previously isolated only a few hundred bacterial species from the body, HMP researchers now calculate that more than 10,000 microbial species occupy the human ecosystem. Moreover, researchers calculate that they have identified between 81 and 99 percent of all microorganismal genera in healthy adults.
“We have defined the boundaries of normal microbial variation in humans,” said James M. Anderson, M.D., Ph.D., director of the NIH Division of Program Coordination, Planning and Strategic Initiatives, which includes the NIH Common Fund. “We now have a very good idea of what is normal for a healthy Western population and are beginning to learn how changes in the microbiome correlate with physiology and disease.”
Microbes that are essential for human survival
HMP researchers also reported that this plethora of microbes contribute more genes responsible for human survival than humans contribute. Where the human genome carries some 22,000 protein-coding genes, researchers estimate that the human microbiome contributes some 8 million unique protein-coding genes or 360 times more bacterial genes than human genes.
This bacterial genomic contribution is critical for human survival. Genes carried by bacteria in the gastro-intestinal tract, for example, allow humans to digest foods and absorb nutrients that otherwise would be unavailable.
“Humans don’t have all the enzymes we need to digest our own diet,” said Lita Proctor, Ph.D., NHGRI’s HMP program manager. “Microbes in the gut break down many of the proteins, lipids and carbohydrates in our diet into nutrients that we can then absorb. Moreover, the microbes produce beneficial compounds, like vitamins and anti-inflammatories that our genome cannot produce.” Anti-inflammatories are compounds that regulate some of the immune system’s response to disease, such as swelling.
Researchers were surprised to discover that the distribution of microbial metabolic activities matters more than the species of microbes providing them. In the healthy gut, for example, there will always be a population of bacteria needed to help digest fats, but it may not always be the same bacterial species carrying out this job.
“It appears that bacteria can pinch hit for each other,” said Curtis Huttenhower, Ph.D., of Harvard School of Public Health and lead co-author for one of the HMP papers in Nature. “It matters whether the metabolic function is present, not which microbial species provides it.”
Moreover, the components of the human microbiome clearly change over time. When a patient is sick or takes antibiotics, the species that makeup of the microbiome may shift substantially as one bacterial species or another is affected. Eventually, however, the microbiome returns to a state of equilibrium, even if the previous composition of bacterial types does not.
Clinical applications
As a part of HMP, NIH funded a number of studies to look for associations of the microbiome with diseases and several PLoS papers include medical results. For example, researchers at the Baylor College of Medicine in Houston compared changes in the vaginal microbiome of 24 pregnant women with 60 women who were not pregnant and found that the vaginal microbiome undergoes a dramatic shift in bacterial species in preparation for birth, principally characterized by decreased species diversity. A newborn is a bacterial sponge as it populates its own microbiome after leaving the sterile womb; passage through the birth canal gives the baby its first dose of microbes, so it may not be surprising that the vaginal microbiome evolved to make it a healthy passage.
Researchers at the Washington University School of Medicine in St. Louis examined the nasal microbiome of children with unexplained fevers, a common problem in children under 3 years of age. Nasal samples from the feverish children contained up to five-fold more viral DNA than children without fever, and the viral DNA was from a wider range of species. Previous studies show that viruses have ideal temperature ranges in which to reproduce. Fevers are part of the body’s defense against pathogenic viruses, so rapid tests for viral load may help children avoid inappropriate treatment with antibiotics that do not kill the viruses but may harm the child’s healthy microbiome.
These are among the earliest clinical studies using microbiome data to study its role in specific illnesses. NIH has funded many more medical studies using HMP data and techniques, including the role of the gut microbiome in Crohn’s disease, ulcerative colitis and esophageal cancer; skin microbiome in psoriasis, atopic dermatitis and immunodeficiency; urogenital microbiome in reproductive and sexual history and circumcision; and a number of childhood disorders, including pediatric abdominal pain, intestinal inflammation, and a severe condition in premature infants in which the intestine actually dies.
“Enabling disease-specific studies is the whole point of the Human Microbiome Project,” said Barbara Methé, Ph.D., of the J. Craig Venter Institute, Rockville, MD, and lead co-author of the Nature paper on the framework for current and future human microbiome research. “Now that we understand what the normal human microbiome looks like, we should be able to understand how changes in the microbiome are associated with, or even cause, illnesses.”
The NIH Common Fund also invested in a series of studies to evaluate the ethical, legal and social implications of microbiome research. While the results of these studies are yet to be published, a number of important issues already have been identified, ranging from how products designed to manipulate the microbiome — such as probiotic concoctions that include live microorganism believed to benefit the body — might be regulated, to whether individuals should begin to consider storing their microbiome while healthy.
Ref.: The Human Microbiome Project Consortium, A framework for human microbiome research, Nature, 2012, DOI: 10.1038/nature11209 (open access)
Ref.: The Human Microbiome Project Consortium, Structure, function and diversity of the healthy human microbiome, Nature, 2012, DOI: 10.1038/nature11234 (open access)
